Gut microbiome and neurocritically ill patients
Article information
Abstract
Since the times of Rokitansky and Cushing, we have been fascinated by the connections between the gut and the brain. Recent advances in next-generation sequencing techniques have shown that this relationship is even more complex and integral to our sense of self than previously imagined. As these techniques refine our understanding of the abundance and diversity of the gut bacterial microbiome, the relationship between the gut and the brain has been redefined. Now, this is understood as a complex symbiotic network with bidirectional communication, the gut-brain axis. The implication of this communication involves an intense focus of research on a variety of chronic psychiatric, neurological, neurodegenerative, and neuro-oncological diseases. Recently, the gut-brain axis has been studied in neurologically ill patients requiring intensive care. Preliminary studies have shown that acute brain injury changes the bacterial phenotype from one that is symbiotic with the host human to one that is pathologic, termed the “pathobiome.” This can contribute to nosocomial pneumonia and sepsis. The first studies in neurologically ill patients in the neurointensive care unit (neuroICU) demonstrated changes in the gut microbiome between neuroICU patients and healthy matched subjects. Specifically, a decrease in short-chain fatty acid-producing bacteria and increase in harmful gut microbes have been associated with mortality and decreased function at discharge. Although these preliminary findings are exciting and have opened a new field of research in the complex neuroICU population, there are several limitations and challenges. Further investigation is needed to confirm these correlations and understand their implications on patients in a complex intensive care environment.
INTRODUCTION
The development of next-generation sequencing technologies has enabled the study of the gut microbiome. The gut microbiome comprises all living organisms that inhabit the human gut. When a human being is healthy, the host and microbiome live symbiotically. In this scenario, the host provides nutrients and an environment for the bacteria to thrive, and the bacteria supply necessary nutrients back to the host [1-3]. This relationship is forged between the brain via the gut-brain axis: a bidirectional communication through fecal metabolites such as neurotransmitters and short-chain fatty acids (SCFAs), the autonomic nervous system (ANS; vagus nerve), and neuroendocrine pathways [4].
Although studies have examined the role of microbiota in a variety of chronic neurological diseases [1-3,5-11], studies on the role of the microbiome in patients with acute brain injury are in their infancy. Recent studies on the gut microbiome in neurocritically ill patients needing care in the neurointensive care unit (neuroICU) have revealed that the gut microbiome is an important factor in disease processes and prognosis of these patients [12-14]. In this review article, we summarize and introduce the concept of the gut microbiome and disease, the gut-brain axis, and its relationship with neurologic diseases in critically ill patients. We highlight the possible implications of the gut-brain axis on patients in the intensive care unit (ICU) and neuroICU and comment on future research areas and their challenges.
GUT MICROBIOME
The gut microbiome, often referred to as the forgotten organ, comprises the microbes that inhabit the gastrointestinal tract. These microbes are more than 10 times as common as the human cells in our bodies and constitute over 150 of the genes of our human genome [4,15]. It is estimated that the adult gut microbiome comprises up to 1,000 species, of which Bacteroidetes and Firmicutes are the two predominant phylotypes [16,17]. The gut microbiome is established early in life and is susceptible to multiple factors, including diet, ethnicity, and age [18-20]. The gut microbiome and its associated genes and products coexist in a homeostatic ecosystem within their host [20].
The functions of the gut microbiome include immune activation and response modulation, epithelial barrier integrity, nutrient absorption and storage, conversion of luminal compounds to metabolites, host-bacterial interactions on the mucosal surface, and long-term behaviors and brain process modulation [5,20-24]. Since the microbiome plays a critical role in the normal physiological function of the gut, multiple studies have pursued the establishment of the taxonomic composition and structure of this microbiome’s constituents in healthy individuals. However, the definition of a “normal” gut microbiome remains inconclusive given the high compositional variability of the microbial taxa, even within healthy individuals and their family members [18,25,26]. However, the genes encoding specific metabolic functions and regulatory pathways are largely conserved [18,20,25,27].
The disruption of the composition and, therefore, the normal function of the microbiome is called dysbiosis. This dysbiosis can be the product of many pathological states, but is usually the product of antibiotics, dietary changes, or a lack of bacterial diversity [20,28]. Dysbiosis is a crucial aspect of the gut microbiome, given that this state increases the host’s susceptibility to the disease owing to its inability to effectively respond to environmental changes [29]. However, whether dysbiosis is a response to or the cause of a particular disease state remains uncertain [20,29].
Microbiome sequencing
The development of two specific techniques, 16S ribosomal RNA (16S rRNA or 16S rRNA) and whole-genome shotgun sequencing (WGS) have enabled us to study the microbiome in greater detail. Both the techniques are similar and provide complementary information. The 16S gene sequencing is mostly used in identifying the microbiome’s bacterial composition. The 16S rRNA has nine variable regions (V1-V9), which distinguish individual bacterial taxa from extracted bacterial DNA. When processed, the DNA sample is amplified by polymerase chain reaction and compared to a known bacterial library to identify the lowest taxonomic levels [30]. While WGS is more expensive and demanding, it provides data on strain-level resolution and functional capacity, characteristics that cannot be obtained with 16S sequencing. In WGS, all DNA in a sample are sequenced using next-generation sequencing. WGS has some advantages over 16S, including the lack of polymerase chain reaction amplification, entire genome sequencing, and strain resolution, which allows greater inference of the gut microbiome [31]. Conversely, the advantages of 16S include a lower cost, the avoidance of host DNA contamination, and the capability to sequence with lower quantities of genetic material [30,31]. After taxonomic assignment, the bacterial composition is frequently evaluated in terms of alpha diversity (within-sample) and beta diversity (between-sample). Alpha diversity is a measure of the species diversity within a sample. This summarizes its richness and uniformity [32]. Beta diversity describes the species diversity between two or more microbial communities in different samples. Bioinformatic tools for visualizing and comparing the diversity and abundance of the microbiome are being developed. Some methodologies can also predict the biological functions of specific microbiome taxa [33].
Metabolomic analysis
Metabolomic analysis, also known as metabolite or metabolomic analysis, refers to the evaluation of metabolites (vitamins, fatty acids, amino acids, and bile acids) produced or regulated by the gut microbiome. In contrast to classic biochemical approaches that evaluate single compounds, metabolomic analysis evaluates a broader series of metabolites to obtain a holistic understanding of the interactions between the microbiome and the condition studied [34]. Different instruments and software exist for metabolomic analysis, depending on the goal of the study, including liquid chromatography with colorimetric array detection, gas chromatography with mass spectrometry, and liquid chromatography-mass spectrometry. The latter is commonly used because of its large biochemical profile in biological samples, especially in gut microbiome studies [34,35]. In gut microbiome metabolomic analysis, the samples are collected, and small molecules are isolated from the sample and analyzed using one of the previously mentioned techniques. After the collection of data, they have to be curated and analyzed using the appropriate software to discover relevant biochemical pathways involved in the gut microbiome brain axis [10].
GUT MICROBIOME BRAIN AXIS
Bidirectional communication between the enteric nervous system and the central nervous system (CNS) forms a network called the gut-brain axis, which is a relatively new but increasingly accepted concept [4]. Gut microbiome dysbiosis was initially studied in diseases related to the gastrointestinal tract, such as irritable bowel syndrome [24]. However, a growing body of evidence reveals that gut microbiome dysbiosis has pathophysiological effects on the CNS [4,24,36]. Preclinical studies have shown that by using gut microbiome manipulation with germ-free, antibiotic-induced depleted, prebiotic/probiotic supplementation, and fecal microbiota transplant animal models, changes in the gut microbiome can alter brain signaling and function including neurotransmitter receptor expression, memory dysfunction, alterations in neuron excitability, and others [36-39]. The translation of these preclinical studies to human populations has been challenging, given the complexity of changes in the human gut microbiome and inter-individual gut microbiome differences. One approach to studying the effects of the gut microbiome on the brain has been to utilize brain imaging to correlate microbial ecology with various neural networks [5,24]. Manipulation of the gut microbiome using antibiotics has shown increased subcortical and frontoparietal brain connectivity, as well as improved cognitive function in a small cohort of minimal hepatic encephalopathic patients [40]. Although preliminary, this suggests that changes in the gut microbiome affect networks in the diseased brain.
The gut microbiome affects neurological function via multiple pathways, including neuroendocrine and immunological pathways, whereas the brain affects the composition of the gut microbiome via the ANS [24]. The ANS has both central and peripheral neurons, which create a brain-gut loop with constant feedback from afferent and efferent fibers. In association with the enteric nervous system, the ANS can induce changes in the gut that affect the gut microbiome, such as gut motility and mucus secretion [5,24]. Most communication depends on the vagus nerve. The vagus nerve afferent neurons provide signals from several gut layers to the nucleus tractus solitaries of the brain, which then act as an emissary of these gut-derived signals to the brain. On the other end, the integrated parasympathetic response is then conducted back through the vagus nerve, producing physical and behavioral changes [5,41]. Sympathetic innervation through less direct pathways primarily serves as the intestinal mucus layer integrity maintenance [42]. The sympathetic ANS is affected by bacterial metabolites in germ-free and antibiotic-treated mice. In this scenario, the SCFA-producing bacteria have a suppressive effect on sympathetic ascending signaling [43].
Another important route through which the gut-brain axis communicates is through gut bacteria-derived metabolites. The gut microbiome regulates metabolite levels by modulating metabolites reactions [23]. SCFA levels have been identified in the cerebrospinal fluid and brain tissue, and these have been associated with numerous CNS diseases [5,10]. SCFAs broadly impact the immune response through the regulation of antigen-presenting cells and production of interleukin-10, TH-1, and TH-17 production [44]. The microbiome also produces and reacts with several neurotransmitters including serotonin, norepinephrine, and other catecholamines. CNS diseases significantly distort fecal neurotransmitter levels outside the physiological range, creating systemic neurotransmitter change [10].
GUT MICROBIOME AND BLOOD-BRAIN BARRIER
The blood-brain barrier (BBB) comprises an endothelial cell tight junction network that contributes to maintaining CNS homeostasis [45]. The BBB prevents the diffusion of pathogens and hydrophilic molecules from the systemic circulation while permitting the passage of critical gases (O2 and CO2) and lipid-soluble substances (glucose) [45]. BBB disruption is a key mechanism of worsening neurological disease in acute neurological diseases, including traumatic brain injury (TBI) and stroke [46,47]. After the initial brain injury, subsequent breakdown of the BBB leads to the propagation of injury, secondary brain injury, and worsening of clinical outcomes. In both preclinical and clinical settings, increased BBB permeability is a marker of disease progression and a therapeutic target [48].
Studies have shown that the gut microbiome influences the development and maintenance of the BBB tight junctions. Germ-free mice (mice without normal gut microbiome flora) have reduced occludin and claudin-5 expression, key BBB tight junction regulators, resulting in increased BBB permeability [49]. These changes started intrauterine and were maintained after birth, demonstrating the importance of the gut microbiome in BBB formation and maintenance. Furthermore, gut colonization of germ-free mice with microbiomes reverses the effects seen in the BBB due to the lack of gut microbiome [49]. Recent studies have further confirmed the relationship between the BBB and the gut microbiome, Wen et al. [50] showed that postoperative cognitive deficits in mice can be aggravated by the administration of antibiotics. Mice that were administered antibiotics showed decreased expression of tight junctions, consequently increasing the BBB permeability. The BBB alterations could be overturned by the administration of Lactobacillus and sodium butyrate [50]. The effects of gut microbiome composition and the function and permeability of the BBB were further tested in adult mammals by Wu et al. [51]. Rhesus monkeys treated with oral amoxycillin-clavulanic acid had increased permeability of the BBB, as measured by the albumin ratio in the cerebrospinal fluid/serum. This was attributed to the detrimental effect of oral antibiotics on acetic acid-and propionic acid-producing microbiome [51]. Although these preclinical findings are exciting given the possibility of BBB modulation in several CNS diseases, their translation remains untested. Fig. 1 shows the gut microbiome and BBB modulation according to the studies presented.
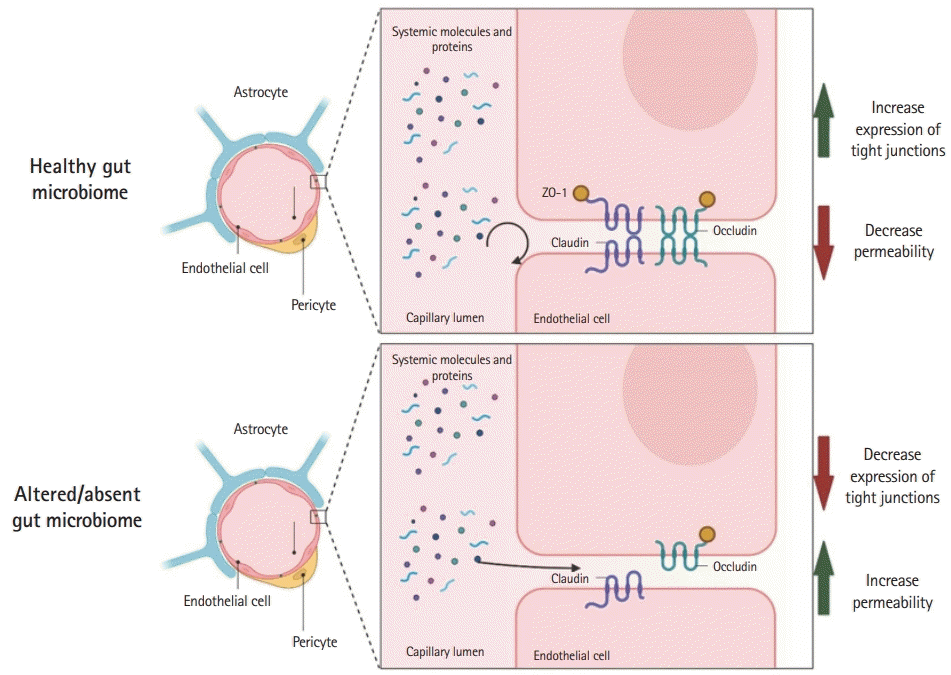
Gut microbiome affects the blood-brain barrier. Animal (mice and monkey) studies have shown that the alteration or absence of the gut microbiome (germ-free or antibiotic-induced) decreased tight junction (occludin and claudin) expression in the blood-brain barrier by an unknown mechanism. This figure was created by authors using BioRender (https://www.BioRender.com).
GUT-BRAIN MICROBIOME AND CNS DISEASES
Accumulating evidence implicates the gut microbiome in various psychiatric, neurological, neurodegenerative, and neuro-oncological diseases [1-3,5-11]. However, the level of evidence varies depending on the disease. Some are still in the preliminary stage, with limited correlational observations, while others provide stronger evidence for a causal role in the disease [5]. The relationship between multiple sclerosis and autism spectrum disorder (ASD) and the gut microbiome has been thoroughly investigated in both animals and humans [3,8,52,53]. In ASD, recent studies have demonstrated that fecal microbiome transplantation is effective in improving gastrointestinal and behavioral symptoms [54]. Moreover, a placebo-controlled study showed that a casein/gluten-free diet combination, along with prebiotic B-GOS, led to behavioral improvement in children with ASD. This was accompanied by a relative increase in the abundance of Bifidobacterium longum [55]. A recent study showed a higher abundance of Akkermansia muciniphila and Acinetobacter calcoaceticus in fecal samples of multiple sclerosis patients [56]. Another recent study showed that A. muciniphila and its associated nicotinamide improved amyotrophic lateral sclerosis symptoms and gene expression patterns, while Ruminococcus torques and Parabacteroides distasonis worsened the amyotrophic lateral sclerosis symptoms [2]. In Alzheimer disease, the gut microbiome is related to β-amyloid plaques and the pathophysiology of the disease [57]. Other interesting findings have correlated epilepsy and Parkinson disease outcomes due to specific microbial-derived metabolites or drug metabolism from bacteria [58,59]. Moreover, psychiatric diseases as well as neuro-oncological entities have been associated with the gut microbiome [5,10,11,60]. In animal models of stroke, several studies have shown the role of the gut microbiome through the modulation of SCFAs [1,61]. Recent studies have shown that the gut microbiome directly impacts the risk of thrombotic events, including strokes, through the production of trimethylamine N-oxide (TMAO), a gut microbe-dependent metabolite produced from precursors known in Western diets (e.g., choline, phosphatidylcholine, carnitine) [62]. TMAO exposure augments intracellular calcium in platelets, with a subsequent increase in thrombosis. This process is dependent on the metabolism of dietary choline and other precursors [62]. Although these studies have shown a strong correlation between multiple CNS diseases and the gut microbiome, much remains to be discovered, especially the causality of the observed changes.
GUT MICROBIOME IN NEUROCRITICAL CARE
The gut microbiome profile of severely ill patients requiring ICU care has only been studied in the last decade. Patients with systemic inflammatory response syndrome with decreased obligate anaerobes and increased pathogenic bacteria showed correlation with septic complications and mortality [63]. Further research has demonstrated that even though the gut microbiome of septic and non-septic critically ill patients is highly heterogeneous, it suffers from low diversity, and is typically colonized by pathogenic microbes (e.g., Enterobacterales, Staphylococcus, Enterococcus, or yeasts) in contrast to a more physiologic gut microbiome [64-66]. This loss of the “normal” gut microbiome generates the absence of important host metabolism functions [65]. Moreover, an increased dysbiosis, measured by the relationship between the two more abundant gut microbiome phyla, Firmicutes and Bacteroides, and the Firmicutes: Bacteroides ratio has been observed in a small prospective cohort of ICU patients [67]. Another potential biomarker of dysbiosis, the gut colonization with Enterococcus [68], has been shown to be closely associated with survival when this bacterium is identified upon admission to the ICU [68,69].
Healthcare-associated infections are one of the many prognostic factors in critically ill patients, including neuroICU patients. This can occur in up to 25% of ICU patients and have significant effects on morbidity and mortality [70]. Rectal or throat colonization by pathogenic bacteria increases the risk of further infection (e.g., pneumonia) with the same pathogen [69,71,72]. In this context, the role of the gut microbiome in the development of pneumonia has been investigated in animal models [73] as well as in the pathogenesis of ICU patients with ventilator-associated pneumonia [74]. Dickson et al. [12] showed that the lung microbiome is enriched with gut microbes in both mouse models and patients with established acute respiratory distress syndrome (ARDS). Additionally, their experiments established that lower gastrointestinal tract bacteria, rather than the upper respiratory tract bacteria, were the culprit of post-sepsis lung infection. Furthermore, Bacteroides were frequently detected in ARDS patients and are associated with the intensity of systemic inflammation [12].
Sepsis, a term used since the time of Hippocrates, has been defined as an infection that provokes life-threatening organ dysfunction [75] and is a common disease worldwide with major repercussions in morbidity, mortality, quality of life, and medical costs for both inpatients and outpatients [76]. The incidence of sepsis is estimated to be 288 per 100,000 person-years in hospital-treated sepsis, whereas hospital mortality is estimated to be 17% for sepsis and 26% for severe sepsis [77]. In the neuroICU, sepsis is a leading cause of morbidity and mortality [78,79]. Although population studies are scarce, some studies estimate it as 1.4%–12.6% of patients admitted to the neuroICU, and it is invariably associated with worsening prognosis [78,79]. A provocative recent study showed that increased bacterial DNA can be identified in the brain after sepsis in both murine models and human patients. These brain-associated bacteria correlate with neuroinflammation and are likely associated with acute brain dysfunction in sepsis [13].
The importance of the gastrointestinal tract in neurological intensive care has been known since the observation of Carl Rokitansky in 1841 and the later classic work of Harvey Cushing regarding the relationship between intracranial pressure and gastric and proximal duodenal ulcers [80,81]. Other known gastrointestinal syndromes that commonly affect neuroICU patients include acute colonic pseudo-obstruction and other motility problems, given the severity of neurological illnesses and the use of medications with gastrointestinal side effects such as opioids and other anticholinergic drugs [82]. Despite the known relationship between neurocritically ill patients and gastrointestinal diseases and the well-known gut microbiome brain axis studies in multiple neurological diseases, little is known about the gut microbiome in the neuroICU population. The first dedicated studies evaluating the microbiome of neurocritically ill patients are the initial steps toward understanding the role of the gut microbiome in the disease process and the mechanisms of neurological patients requiring ICU treatment. This understanding is the first step toward the identification of possible microbiome modulation that affects prognosis. Xu et al. [14] observed a distinct gut microbiome between neuroICU patients and healthy controls. Alpha diversity and the abundance of known SCFA producer bacteria like Ruminococcaceae and Lachnospiraceae were significantly reduced during the hospital stay in neurocritically ill patients. Similar to other studies on critically ill patients [68], Xu et al. [14] showed that neuroICU patients with increased Enterobacteriaceae (family Enterococcus) during their first week of hospital stay had increased 6-month mortality after adjustment for multiple factors. Moreover, an association between Enterobacteriaceae and the modified Rankin scale score at discharge was noted [14]. Although this study represents an important effort to understand the relationship between the gut microbiome and outcomes in neuroICU patients, it has several limitations, including the lack of data regarding antibiotic utilization as well as the heterogeneity of the patient population. A summary of the gut-brain axis relationship in brain injury is shown in Fig. 2.
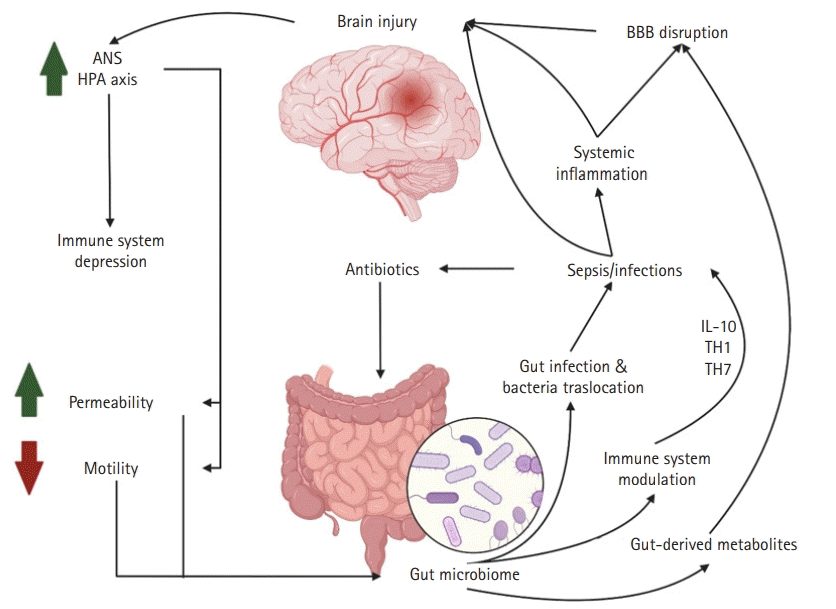
Gut-brain axis in neurocritically ill patients. We hypothesize that patients with severe brain injury (e.g., traumatic brain injury or intraparenchymal hemorrhage), may have dysregulation of the autonomic nervous system (ANS) and hypothalamic-pituitary-adrenal (HPA) axis which could lead to immune system depression, increased gut permeability, and decreased motility. These changes result in gut microbiome dysbiosis, which facilitates infection and bacterial gut translocation, as well as immunosuppression, thereby predisposing individuals to infections and sepsis. An infectious state produces systemic inflammation that promotes blood-brain barrier (BBB) disruption and additional brain injury, secondary to inflammation. Additionally, antibiotic treatment against previously mentioned infections promotes gut dysbiosis. Another mechanism of BBB disruption leading to additional brain injury are gut-derived metabolite changes that are secondary to gut microbiome dysbiosis. IL-10, interleukin-10. This figure was created by authors using BioRender (https://www.BioRender.com).
FUTURE DIRECTION AND CHALLENGES
The gut microbiome in neuroICU patients is an emerging topic of research that requires further investigation. Since research efforts have been performed to elucidate the gut microbiome relationship in multiple CNS diseases (e.g., stroke, brain tumors, epilepsy, or TBI) and a growing body of evidence has shown that the gut microbiome influences common neuroICU comorbidities such as ventilator associated pneumonia and sepsis, we foresee that this topic will become more relevant with the continuity of research efforts. To date, the few published studies have been limited to assessing correlations, for example, Enterobacteria and pneumonia/sepsis; however, future studies should seek to answer more questions pertaining to causality, such as whether the eradication of the pathobiome or restoration of a healthy microbiome through gut manipulation (probiotics, prebiotics, fecal transplants) improves morbidity or mortality. Addressing these questions can open innovative therapeutic avenues for neurocritically ill patients, as established in other diseases such as Clostridium difficile infections [83], or enhance known therapies, as shown in the first human clinical trials of fecal transplant to overcome anti-PD-1 (programmed death-ligand 1) resistance in melanoma [84,85]. These and other gut microbiome therapeutic strategies have revolutionized our understanding of our interactions with the microbiome that inhabits us.
Some challenges of studying the gut microbiome in neuroICU patients are common to gut microbiome studies, such as the lack of a “normal” gut microbiome and differences in the gut microbiome associated with diet and customs. However, other challenges are specific to the neuroICU population, which include a heterogeneous population (TBI, stroke, brain tumors, subarachnoid hemorrhages), high usage of antibiotics, different ICU biomes, and the difficulty of assessing whether the microbiome changes are due to the disease itself, the ICU environment, or medications (Fig. 3).
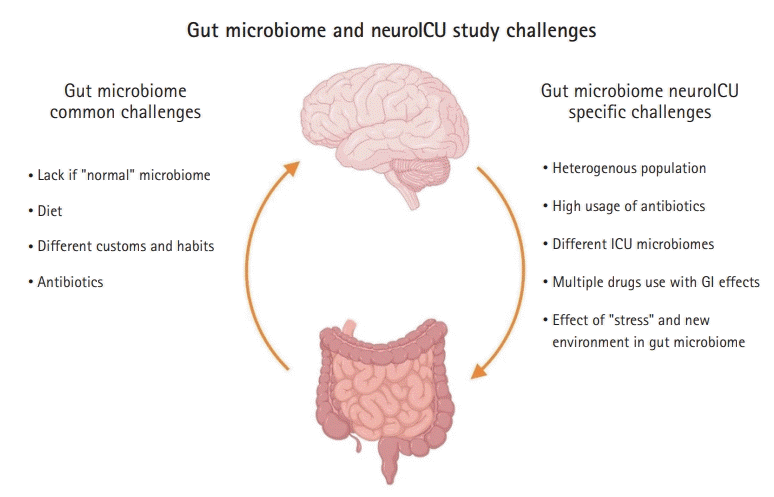
Challenges of gut microbiome studies in the neurointensive care unit (neuroICU). ICU, intensive care unit; GI, gastrointestinal. This figure was created by authors using BioRender (https://www.BioRender.com).
CONCLUSIONS
The gut-brain axis is a bidirectional communication through several pathways, a symbiotic relationship that can be affected by both microbiome changes to pathogens and CNS diseases. The role of the gut microbiome in CNS diseases has been actively studied, but remains an area of limited knowledge.
Notes
Ethics statement
Not applicable.
Conflict of interest
Huimahn A. Choi is an editorial board member of the journal but he was not involved in the peer reviewer selection, evaluation, or decision process of this article. No other potential conflicts of interest relevant to this article were reported.
Acknowledgments
Yoshua Esquenazi is supported by the Center for Clinical and Translational Sciences, which is funded by the National Institutes of Health Clinical and Translational Award UL1 TR003167 from the National Center for Advancing Translational Sciences. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author contributions
Conceptualization: HAC. Methodology: all authors. Project administration: HAC. Visualization: AD. Writing–original draft: AD, HAC. Writing–review & editing: all authors.
